LFF_spatial_ERC
LFF Spatial ERC
Overview
Welcome to the LFF_spatial_ERC project! 🧠
In this work we built a transcriptional profile of the human entorhinal cortex (ERC) with single nucleus RNA sequencing (snRNA-seq) and spatially resolved transcriptomics (SRT) from 31 neurotypical post-mortem donors with diverse risk for Alzheimer’s disease (AD). SRT data was generated with 10x Genomics Visium, snRNA-seq was generated with 10x Genomics Chromium.
We then assessed changes in gene expression associated with risk of AD by APOE carrier status (E2+ vs. E4+), as well as APOE changes specific to ancestry groups (African ancestry or European ancestry), and sex.
This work is being was performed by members of Leonardo Collado-Torres, Kristen Maynard, and Keri Martinowich, teams at the Lieber Institute for Brain Development as well as Mina Ryten’s group from UK DRI Cambridge.
Thank you for your interest in our work!
Study Design
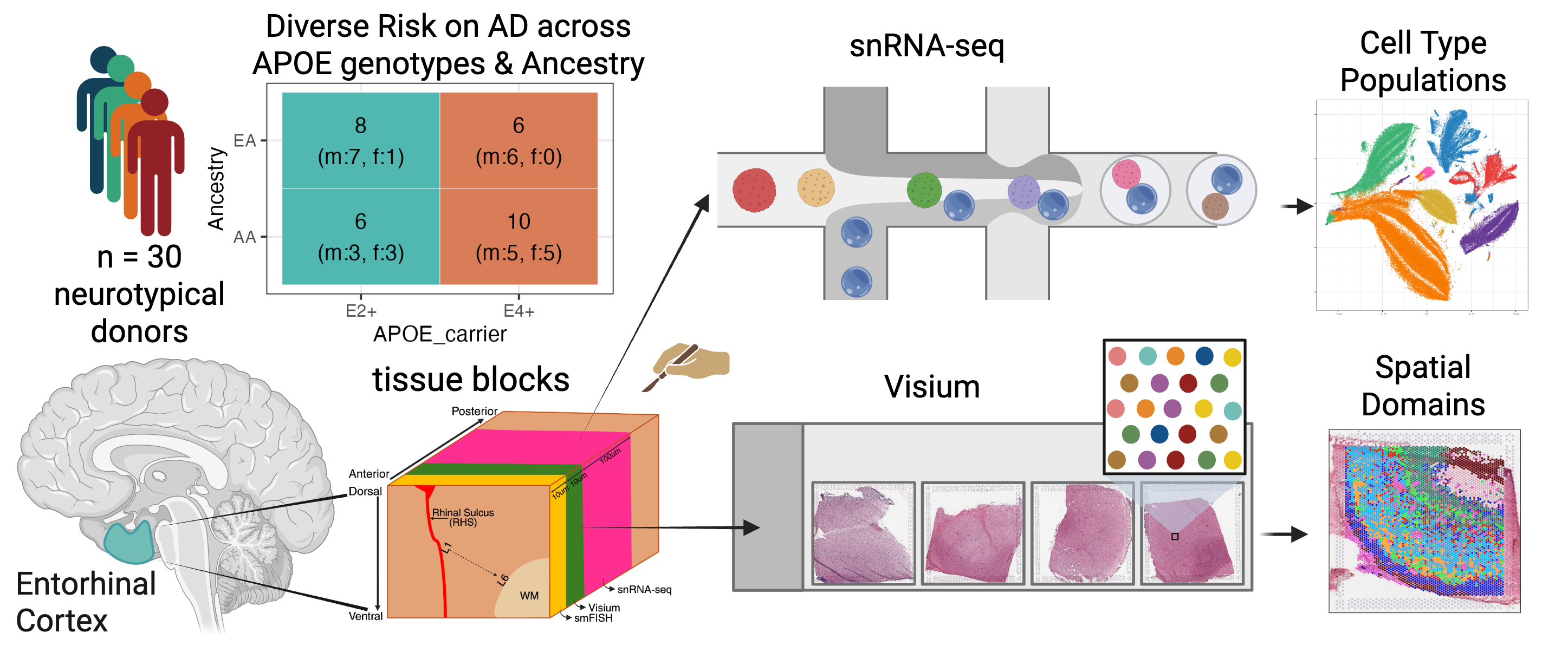
Schematic depicting cohort composition stratified across APOE genotype, sex, and ancestry as well as experimental design for generation of snRNA-seq and SRT data from postmortem human ERC tissue.
Interactive Websites
We provide the following interactive websites to explore the ERC data:
-
🔭 LFF_ERC_Visium: Visium data for 31 samples post-QC w/ clustering and modeling results
-
🔭 LFF_ERC_Visium_QC: Visium data for 31 samples pre-QC drops (including out-tissue spots)
-
👀 iSEE: snRNA-seq data for 31 samples
All of these interactive websites are powered by open source software, namely:
- 🔭
spatialLIBDSee spatialLIBD website for more information - 👀
iSEE
Citing our work
Please cite our pre-print 10.1101/2025.11.20.689483 if you use data from this project.
Huuki-Myers, L. A., Divecha, H. R., Bach, S. V., Valentine, M. R., Eagles, N. J., Mulvey, B., Bharadwaj, R. A., Zhang, R., Evans, J. R., Grant-Peters, M., Miller, R. A., Kleinman, J. E., Han, S., Hyde, T. M., Page, S. C., Weinberger, D. R., Martinowich, K., Ryten, M., Maynard, K. R. & Collado-Torres, L. apoe E4 alzheimer’s risk converges on an oligodendrocyte subtype in the human entorhinal cortex. BioRxiv (2025). doi:10.1101/2025.11.20.689483
Below is the citation in BibTeX format.
@article{Huuki-Myers_Divecha_Bach_Valentine_Eagles_Mulvey_Bharadwaj_Zhang_Evans_Grant-Peters_et al._2025,
title={apoe E4 alzheimer’s risk converges on an oligodendrocyte subtype in the human entorhinal cortex},
DOI={10.1101/2025.11.20.689483},
abstractNote={<p>The entorhinal cortex (ERC) is implicated in early
progression of Alzheimer’s disease (AD). Here we investigated the impact of
established biological risk factors for AD, including APOE genotype
(E2 versus E4 alleles), sex, and ancestry, on gene expression in the human
ERC. We generated paired spatially-resolved transcriptomics (SRT) and
single-nucleus RNA sequencing data (snRNA-seq) in postmortem human ERC
tissue from middle aged brain donors with no history of AD. APOE-dependent
changes in gene expression predominantly mapped to a
transcriptionally-defined oligodendrocyte subtype, which varied
substantially with ancestry, and suggested differences in oligodendrocyte
differentiation and myelination. Integration of SRT and snRNA-seq data
identified a common gene expression signature associated with APOE genotype,
which we localized to the same oligodendrocyte subtype and a white matter
spatial domain. This suggests that AD risk in ERC may be associated with
disrupted oligodendrocyte function, potentially contributing to future
neurodegeneration.</p>},
journal={BioRxiv},
author={Huuki-Myers, Louise A. and Divecha, Heena R. and Bach, Svitlana V.
and Valentine, Madeline R. and Eagles, Nicholas J. and Mulvey, Bernard and
Bharadwaj, Rahul A. and Zhang, Ruth and Evans, James R. and Grant-Peters,
Melissa and et al.},
year={2025},
month={Nov}}
Data & Code avalibility
Data and code for this project are available on Github.
Organization of code, data, and plots follows our team’s project template.
Raw data generated as a part of this study have been deposited on the Gene Expression Omnibus (GEO) with accession numbers GSE307990 (SRT data) and GSE308007 (snRNA-seq data).
Processed data files (R objects) used to make the interactive apps (logcounts only)
can be found on our Globus Endpoints under the
heading jhpce#LFF_ERC.
R objects with both the counts and logcounts can be downloaded through the fetch_data() function from spatialLIBD website v1.23.1 or newer.
Access data with spatialLIBD
## SRT data
spe_erc_path <- spatialLIBD::fetch_data(type = "LFF_spatial_ERC_SRT")
spe_erc_path <- unzip(spe_erc_path, exdir = tempdir())
spe_erc <- HDF5Array::loadHDF5SummarizedExperiment(
file.path(tempdir(), "spe_ERC_annotated")
)
spe_erc
# dim: 30494 122202
# lobstr::obj_size(spe_erc) 3.20 GB
## SCE data
sce_erc_path <- spatialLIBD::fetch_data(type = "LFF_spatial_ERC_snRNAseq")
sce_erc_path <- unzip(sce_erc_path, exdir = tempdir())
sce_erc <- HDF5Array::loadHDF5SummarizedExperiment(
file.path(tempdir(), "sce_ERC_subcluster")
)
sce_erc
# dim: 38606 122004
# lobstr::obj_size(sce_erc) 258.20 MB
Source code is under the MIT License and permanently archived on Zenodo. The GitHub repository includes log files with specific software versions used for each analysis.
Installation and Requirements
All software versions are listed in log files. The R session information was
automatically generated with sessioninfo::session_info().
The code repo can be downloaded via git clone to a normal desktop, this may
take up to an hour given it’s size.
NOTE this code is specialized for this project’s data, and will need to be adapted to run on other datasets.
Funding
This project was supported thanks to the Carol and Gene Ludwig Award for Neurodegeneration Research (DRW) from the Ludwig Famiy Foundation and the Lieber Institute for Brain Development.
Main Data
snRNA-seq
SingleCellExperiment
- project path:
"processed-data/sce_objects/sce_ERC_subcluster"(Not on github due to large file size: 36G)
## Load HD5F sce
sce <- HDF5Array::loadHDF5SummarizedExperiment(here::here("processed-data", "sce_objects", "sce_ERC_subcluster"))
# class: SingleCellExperiment
# dim: 38606 122004
Input files
-
FASTQ:
raw-data/FASTQ_snRNAseq -
Sample ID table:
processed-data/04_snRNA-seq/erc_sn_sample_info.csv
Pseudobulked snRNA-seq & modeling data
-
cell type subcluster pseudobulk:
processed-data/04_snRNA-seq/29_sn_subcluster_model_pseudobulk/sce_subcluster_pseudobulk-cell_type_anno.rds -
cell type subcluster modeling:
processed-data/04_snRNA-seq/29_sn_subcluster_model_pseudobulk/sce_subcluster_modeling_results-cell_type_anno.rds" -
cell type broad pseudobulk:
processed-data/04_snRNA-seq/29_sn_subcluster_model_pseudobulk/sce_subcluster_pseudobulk-cell_type_broad.rds -
cell type broad modeling:
processed-data/04_snRNA-seq/29_sn_subcluster_model_pseudobulk/sce_subcluster_modeling_results-cell_type_broad.rds
SRT/Visium
SpatialExperiment
-
project path:
"processed-data/spe_objects/spe_ERC_annotated" -
pre-QC version:
processed-data/spe_objects/spe_raw.rds(Not on github due to large file size: 4G)
## Load HD5F spe
spe <- HDF5Array::loadHDF5SummarizedExperiment(here::here("processed-data", "spe_objects", "spe_ERC_annotated"))
# class: SpatialExperiment
# dim: 30494 122202
Input files
-
FASTQ:
raw-data/FASTQ -
Images:
raw-data/FASTQ/Images -
SAMPLE ID table:
processed-data/02_build_spe/sample_info.csv
Pseudobulked Visium & Modeling data
-
pseudobulk:
processed-data/05_spe_correct_cluster/20_model_pseudobulk_anno/spe_pseudobulk-SpD.rds -
modeling data:
processed-data/05_spe_correct_cluster/20_model_pseudobulk_anno/modeling_results-SpD.rds
Internal
JHPCE location: /dcs05/lieber/marmaypag/LFF_spatialERC_LIBD4140/LFF_spatial_ERC/
snRNA-seq main data JHPCE path: "/dcs05/lieber/marmaypag/LFF_spatialERC_LIBD4140/LFF_spatial_ERC/processed-data/sce_objects/sce_ERC_subcluster"
SRT main data JHPCE path: /dcs05/lieber/marmaypag/LFF_spatialERC_LIBD4140/LFF_spatial_ERC/processed-data/spe_objects/spe_ERC_annotated