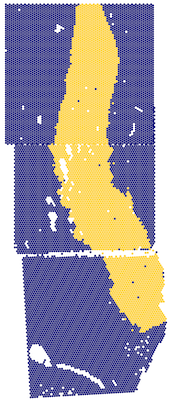
Add transformed array and pixel coordinates to a SpatialExperiment
Source: R/add_array_coords.R
add_array_coords.RdGiven a SpatialExperiment-class, sample information, and coordinates produced from the refinement workflow, add array and pixel coordinates appropriate for the linearly transformed capture areas making up each group present in the SpatialExperiment-class.
Usage
add_array_coords(
spe,
sample_info,
coords_dir,
calc_error_metrics = FALSE,
algorithm = c("LSAP", "Euclidean")
)Arguments
- spe
A SpatialExperiment-class object.
- sample_info
A
data.frame()with columnscapture_area,group,fiji_xml_path,fiji_image_path,spaceranger_dir,intra_group_scalar, andgroup_hires_scalef. The last two are made byrescale_fiji_inputs().- coords_dir
A
character(1)vector giving the directory containing sample directories each withtissue_positions.csv,scalefactors_json.json, andtissue_lowres_image.pngfiles produced from refinement with prep_fiji_coords() and related functions.- calc_error_metrics
A
logical(1)vector indicating whether to calculate error metrics related to mapping spots to well-defined array coordinates. IfTRUE, addseuclidean_errorandshared_neighborsspot-level metrics to thecolData(). The former indicates distance in number of inter-spot distances to "move" a spot to the new array position; the latter indicates the fraction of neighbors for the associated capture area that are retained after mapping, which can be quite time-consuming to compute.- algorithm
A
character(1)vector indicating which mapping algorithm to employ when computing group-wide array coordinates. The default of "LSAP" is generally recommended, as it guarantees one-to-one mappings at the cost of computational time and some Euclidean error. The faster alternative "Euclidean" minimizes Euclidean error but may produce duplicate mappings, which is generally undesirable downstream (for clustering, etc).
Value
A SpatialExperiment-class
object with additional colData
columns pxl_row_in_fullres_[suffix] and pxl_col_in_fullres_[suffix]
with [suffix] values original and rounded;
array_row_original and array_col_original columns; and
modified colData() columns array_row and
array_col and spatialCoords() with their transformed values.
Details
Array coordinates are determined via an algorithm that fits each spot to
the nearest spot on a new, imaginary, Visium-like capture area. The imaginary
capture area differs from a real capture area only in its extent; array
coordinates still start at 0 but may extend arbitrarily beyond the normal
maximum indices of 77 and 127 to fit every capture area in each group
defined in the SpatialExperiment-class.
The goal is to return well-defined
array coordinates in a consistent spatial orientation for each group, such
that downstream applications, such as clustering with BayesSpace, can
process each group as if it really were one capture area in the first place.
See
https://research.libd.org/visiumStitched/articles/visiumStitched.html#defining-array-coordinates
for more details.
Examples
if (!exists("spe")) {
spe <- spatialLIBD::fetch_data(type = "visiumStitched_brain_spe")
}
#> 2025-10-13 17:51:35.607308 loading file /github/home/.cache/R/BiocFileCache/201e1dbfb001_visiumStitched_brain_spe.rds%3Frlkey%3Dnq6a82u23xuu9hohr86oodwdi%26dl%3D1
########################################################################
# Prepare sample_info
########################################################################
sample_info <- dplyr::tibble(
group = "Br2719",
capture_area = c("V13B23-283_A1", "V13B23-283_C1", "V13B23-283_D1")
)
# Add 'spaceranger_dir' column
sr_dir <- tempdir()
temp <- unzip(
spatialLIBD::fetch_data("visiumStitched_brain_spaceranger"),
exdir = sr_dir
)
#> 2025-10-13 17:51:40.172261 loading file /github/home/.cache/R/BiocFileCache/201ee55b2c_visiumStitched_brain_spaceranger.zip%3Frlkey%3Dbdgjc6mgy1ierdad6h6v5g29c%26dl%3D1
sample_info$spaceranger_dir <- file.path(
sr_dir, sample_info$capture_area, "outs", "spatial"
)
# Add Fiji-output-related columns
fiji_dir <- tempdir()
temp <- unzip(
spatialLIBD::fetch_data("visiumStitched_brain_Fiji_out"),
exdir = fiji_dir
)
#> 2025-10-13 17:51:42.077507 loading file /github/home/.cache/R/BiocFileCache/201e7712fb23_visiumStitched_brain_fiji_out.zip%3Frlkey%3Dptwal8f5zxakzejwd0oqw0lhj%26dl%3D1
sample_info$fiji_xml_path <- temp[grep("xml$", temp)]
sample_info$fiji_image_path <- temp[grep("png$", temp)]
## Re-size images and add more information to the sample_info
sample_info <- rescale_fiji_inputs(sample_info, out_dir = tempdir())
## Preparing Fiji coordinates and images for build_SpatialExperiment()
spe_input_dir <- tempdir()
prep_fiji_coords(sample_info, out_dir = spe_input_dir)
#> [1] "/tmp/RtmpDl3GYr/Br2719/tissue_positions.csv"
prep_fiji_image(sample_info, out_dir = spe_input_dir)
#> [1] "/tmp/RtmpDl3GYr/Br2719/tissue_lowres_image.png"
#> [2] "/tmp/RtmpDl3GYr/Br2719/scalefactors_json.json"
########################################################################
# Add array coordinates
########################################################################
# Run with Euclidean algorithm for speed. On real analyses, "LSAP" is
# generally recommended.
spe_new <- add_array_coords(
spe, sample_info, tempdir(), algorithm = "Euclidean"
)
# Several columns related to spatial coordinates were added
added_cols_regex <- "^(array|pxl)_(row|col)(_in_fullres)?_(original|rounded)$"
colnames(SummarizedExperiment::colData(spe_new))[
grep(added_cols_regex, colnames(SummarizedExperiment::colData(spe_new)))
]
#> [1] "array_row_original" "array_col_original"
#> [3] "pxl_col_in_fullres_rounded" "pxl_row_in_fullres_rounded"
#> [5] "pxl_row_in_fullres_original" "pxl_col_in_fullres_original"
# 'array_row', 'array_col', and spatialCoords() were overwritten with
# their transformed values
head(spe$array_row)
#> [1] 0 50 3 59 14 43
head(spe$array_col)
#> [1] 16 102 43 19 94 9
head(SpatialExperiment::spatialCoords(spe_new))
#> pxl_col_in_fullres pxl_row_in_fullres
#> AAACAACGAATAGTTC-1 1874 50718
#> AAACAAGTATCTCCCA-1 13935 38805
#> AAACAATCTACTAGCA-1 2599 46977
#> AAACACCAATAACTGC-1 16101 50308
#> AAACAGAGCGACTCCT-1 5255 39910
#> AAACAGCTTTCAGAAG-1 12242 51692