Introduction to smokingMouse
Daianna Gonzalez-Padilla
Lieber Institute for Brain Development (LIBD)glezdaianna@gmail.com
31 March 2026
Source:vignettes/smokingMouse.Rmd
smokingMouse.RmdIntroduction
Welcome to the smokingMouse project. In this vignette
we’ll show you how to access the smoking-nicotine-mouse LIBD datasets.
You can find the analysis code and the data generation in here.
Motivation
The main motivation to create this Bioconductor package was to provide public and free access to all RNA-seq datasets that were generated for the smoking-nicotine-mouse project, containing variables of interest that make it possible to answer a wide range of biological questions related to smoking and nicotine effects in mice.
Overview
This bulk RNA-sequencing project consisted of a differential expression analysis (DEA) involving 4 data types: genes, transcripts, exons, and exon-exon junctions. The main goal of this study was to explore the effects of prenatal exposure to smoking and nicotine on the developing mouse brain. As secondary objectives, this work evaluated: 1) the affected genes by each exposure in the adult female brain in order to compare offspring and adult results, and 2) the effects of smoking on adult blood and brain to search for overlapping biomarkers in both tissues. Finally, DEGs identified in mice were compared against previously published results in human (Semick et al., 2020 and Toikumo et al., 2023).
Study design
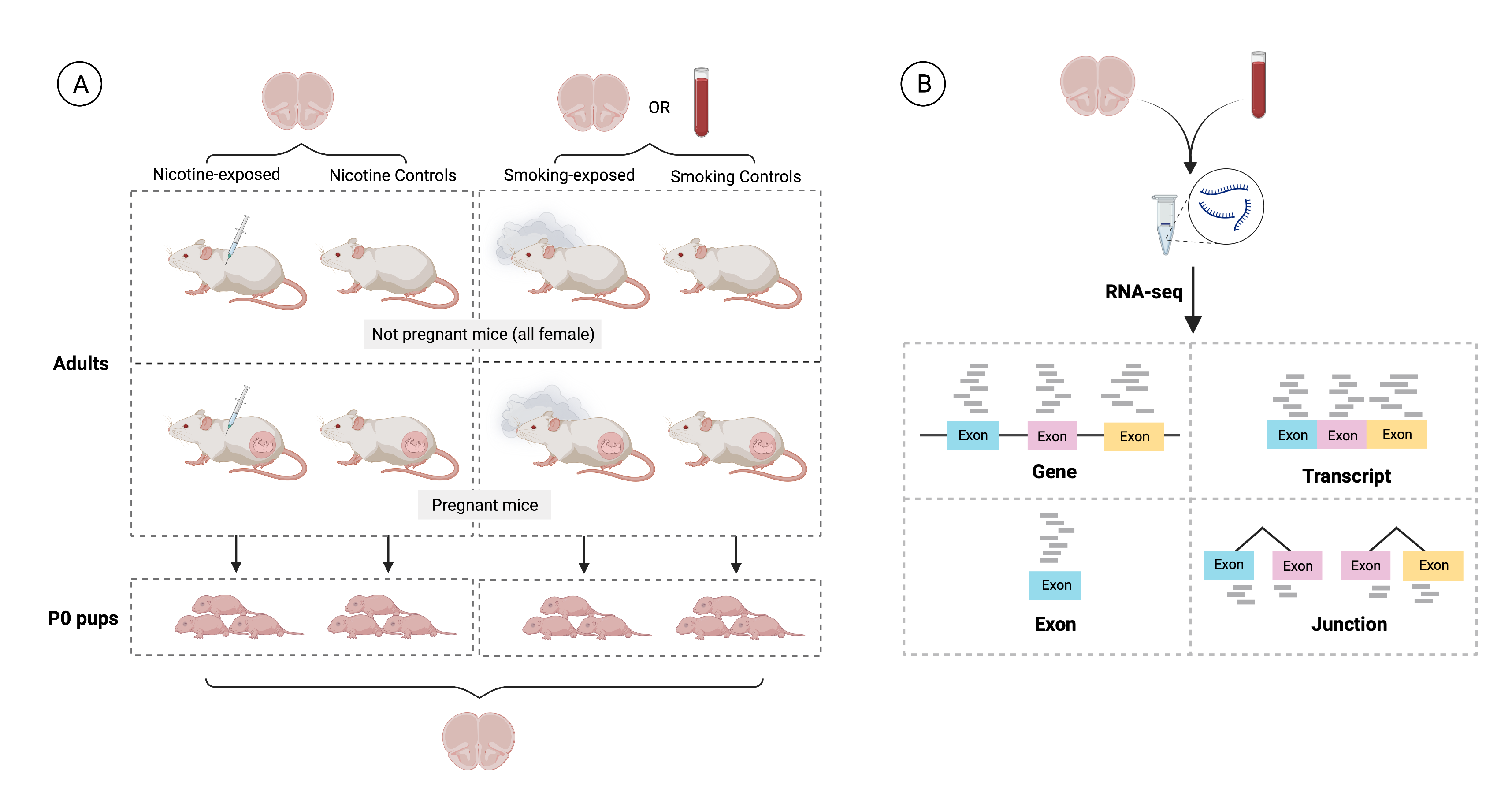
Experimental design of the study. A) 21 pregnant mice were split into two experiments: in the first one prenatal nicotine exposure (PNE) was modeled administering nicotine (n=3) or vehicle (n=3) to the dams during gestation, and in the second maternal smoking during pregnancy (MSDP) was modeled exposing dams to cigarette smoke during gestation (n=8) or using them as controls (n=7). A total of 137 pups were born: 19 were born to nicotine-administered mice, 23 to vehicle-administered mice, 46 to smoking-exposed mice, and 49 to smoking control mice. 17 nonpregnant adult females were also nicotine-administered (n=9) or vehicle-administered (n=8) to model adult nicotine exposure, and 9 additional nonpregnant dams were smoking-exposed (n=4) or controls (n=5) to model adult smoking. Frontal cortex samples of all P0 pups (n=137: 42 for PNE and 95 for MSDP) and adults (n=47: 23 for the nicotine experiment and 24 for the smoking experiment) were obtained, as well as blood samples from the smoking-exposed and smoking control adults (n=24), totaling 208 samples. Number of donors and samples are indicated in the figure. B) RNA was extracted from such samples and bulk RNA-seq experiments were performed, obtaining expression counts for genes, transcripts, exons, and exon-exon junctions.
Workflow
The next table summarizes the analyses done at each feature level.
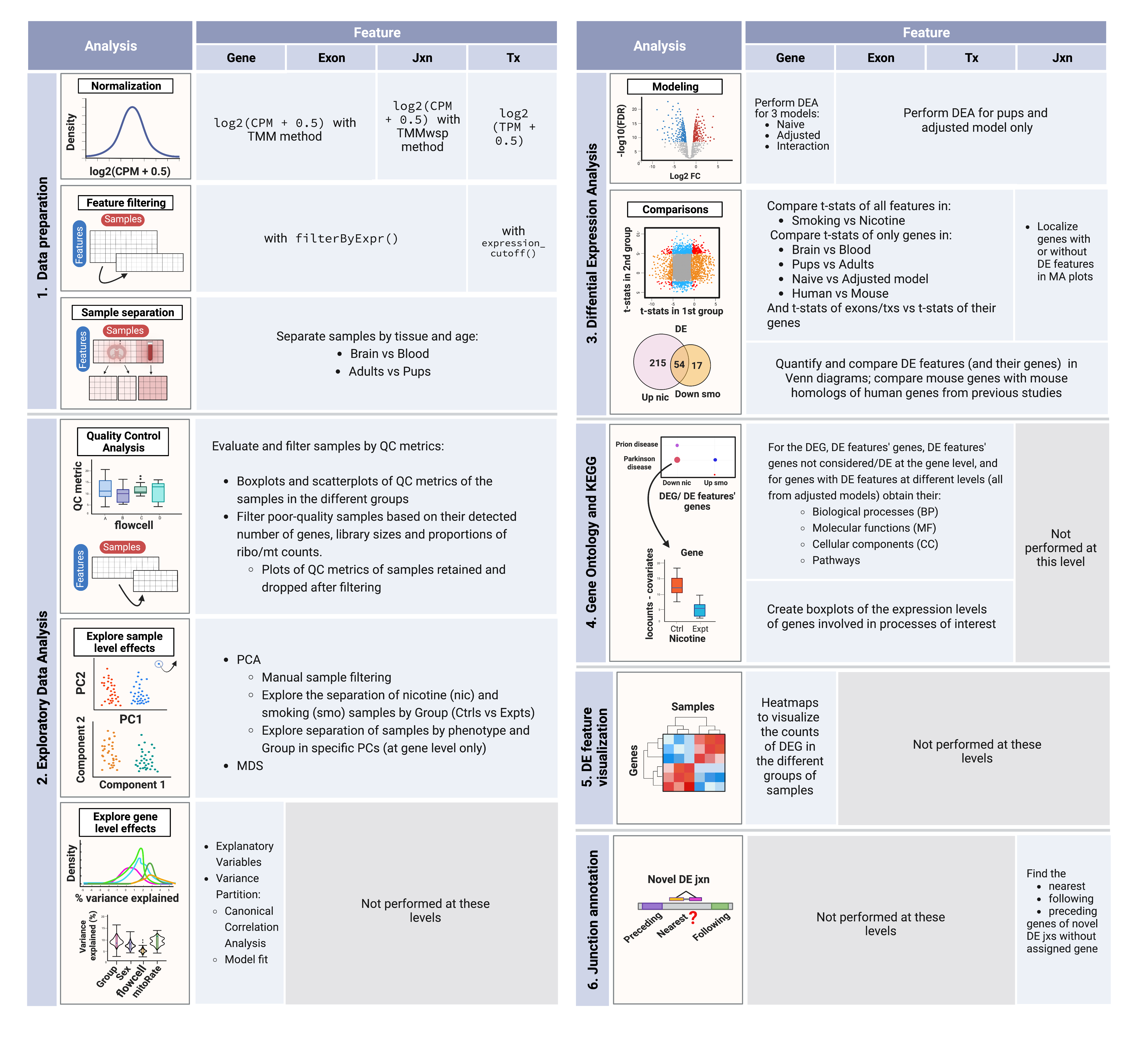
Summary of analysis steps across gene expression feature levels:
1. Data processing: counts of genes, exons, and exon-exon junctions were normalized to CPM and log2-transformed; transcript expression values were only log2-transformed since they were already in TPM. Lowly-expressed features were removed using the indicated functions and samples were separated by tissue and age in order to create subsets of the data for downstream analyses.
2. Exploratory Data Analysis (EDA): QC metrics of the samples were examined and used to filter the poor quality ones. Sample level effects were explored through dimensionality reduction methods and segregated samples in PCA plots were removed from the datasets. Gene level effects were evaluated with analyses of variance partition.
3. Differential Expression Analysis (DEA): with the relevant variables identified in the previous steps, the DEA was performed at the gene level for nicotine and smoking exposure in adult and pup brain samples, and for smoking exposure in adult blood samples; DEA at the rest of the levels was performed for both exposures in pup brain only. DE signals of the genes in the different conditions, ages, tissues, and species (using human results from Semick et al., 2020) were contrasted, as well as the DE signals of exons and transcripts vs those of their genes. Mean expression of DEGs and non-DEGs genes with and without DE features was also analyzed. Then, all resultant DEGs and DE features (and their genes) were compared by direction of regulation (up or down) between and within exposures (nicotine/smoking); mouse DEGs were also compared against human genes associated with TUD from Toikumo et al., 2023.
4. Functional Enrichment Analysis: GO & KEGG terms significantly enriched in the clusters of DEGs and genes of DE transcripts and exons were obtained.
5. DGE visualization: the log2-normalized expression of DEGs was represented in heat maps in order to distinguish the groups of up- and down-regulated genes.
6. Novel junction gene annotation: for uncharacterized DE junctions with no annotated gene, their nearest, preceding, and following genes were determined.
Abbreviations: Jxn: junction; Tx(s): transcript(s); CPM: counts per million; TPM: transcripts per million; TMM: Trimmed Mean of M-Values; TMMwsp: TMM with singleton pairing; QC: quality control; PC: principal component; DEA: differential expression analysis; DE: differential expression/differentially expressed; FC: fold-change; FDR: false discovery rate; DEGs: differentially expressed genes; TUD: tobacco use disorder; DGE: differential gene expression.
All R scripts created to perform such analyses can be
found in code
on GitHub.
Basics
Install smokingMouse
R is an open-source statistical environment which can be
easily modified to enhance its functionality via packages. smokingMouse
is an R package available via the Bioconductor repository for packages.
R can be installed on any operating system from CRAN after which you can install
smokingMouse
by using the following commands in your R session:
if (!requireNamespace("BiocManager", quietly = TRUE)) {
install.packages("BiocManager")
}
BiocManager::install("smokingMouse")
## Check that you have a valid Bioconductor installation
BiocManager::valid()Required knowledge
smokingMouse is based on many other packages, in particular those that have implemented the infrastructure needed for dealing with RNA-seq data and differential expression. That is, packages such as SummarizedExperiment and limma.
If you are asking yourself the question “Where do I start using Bioconductor?” you might be interested in this blog post.
Asking for help
As package developers, we try to explain clearly how to use our
packages and in which order to use the functions. But R and
Bioconductor have a steep learning curve so it is critical
to learn where to ask for help. The blog post quoted above mentions some
but we would like to highlight the Bioconductor support site
as the main resource for getting help: remember to use the
smokingMouse tag and check the older
posts. Other alternatives are available such as creating GitHub
issues and tweeting. However, please note that if you want to receive
help you should adhere to the posting
guidelines. It is particularly critical that you provide a small
reproducible example and your session information so package developers
can track down the source of the error.
Citing smokingMouse
We hope that smokingMouse will be useful for your research. Please use the following information to cite the package and the overall approach.
## Citation info
citation("smokingMouse")
#> To cite package 'smokingMouse' in publications use:
#>
#> Gonzalez-Padilla D, Collado-Torres L (2026). _Provides access to
#> smokingMouse project data_. doi:10.18129/B9.bioc.smokingMouse
#> <https://doi.org/10.18129/B9.bioc.smokingMouse>.
#> https://github.com/LieberInstitute/smokingMouse/smokingMouse - R
#> package version 1.9.0,
#> <http://www.bioconductor.org/packages/smokingMouse>.
#>
#> Gonzalez-Padilla D, Eagles NJ, Cano M, Pertea G, Jaffe AE, Maynard
#> KR, Hancock DB, Handa JT, Martinowich K, Maynard KR, Collado-Torres L
#> (2024). "Molecular impact of nicotine and smoking exposure on the
#> developing and adult mouse brain." _bioRxiv_.
#> doi:10.1101/2024.11.05.622149
#> <https://doi.org/10.1101/2024.11.05.622149>.
#> <https://doi.org/10.1101/2024.11.05.622149>.
#>
#> To see these entries in BibTeX format, use 'print(<citation>,
#> bibtex=TRUE)', 'toBibtex(.)', or set
#> 'options(citation.bibtex.max=999)'.Quick start to using smokingMouse
To get started, please load the smokingMouse package.
smokingMouse datasets
The raw expression data were generated by LIBD researchers and are composed of read counts of genes, exons, and exon-exon junctions (jxns), and transcripts-per-million (TPM) of transcripts (txs), across the 208 mice samples (from brain/blood; adults/pups; nicotine-exposed/smoking-exposed/controls). The datasets available in this package were generated by Daianna Gonzalez-Padilla. The human data were generated in Semick et al., 2018 and contain the results of a DEA in adult and prenatal human brain samples exposed to cigarette smoke.
Description of the datasets
Mouse datasets:
- They are 4 RangedSummarizedExperiment
(RSE) objects that contain feature info in
rowData(RSE)and sample info incolData(RSE). - Raw expression counts (and TPM for txs) can be accessed with
assays(RSE)$countsand the log-transformed data (log2(CPM + 0.5) for genes, exons, and jxns, and log2(TPM + 0.5) for txs) withassays(RSE)$logcounts.
Data specifics
-
‘rse_gene_mouse_RNAseq_nic-smo.Rdata’:
(
rse_geneobject) the gene RSE object contains the raw and log-normalized expression data of 55,401 mouse genes across the 208 samples from brain and blood of control and nicotine/smoking-exposed pup and adult mice. -
‘rse_tx_mouse_RNAseq_nic-smo.Rdata’: (
rse_txobject) the tx RSE object contains the raw and log-scaled expression data of 142,604 mouse transcripts across the 208 samples from brain and blood of control and nicotine/smoking-exposed pup and adult mice. -
‘rse_exon_mouse_RNAseq_nic-smo.Rdata’:
(
rse_exonobject) the exon RSE object contains the raw and log-normalized expression data of 447,670 mouse exons across the 208 samples from brain and blood of control and nicotine/smoking-exposed pup and adult mice. -
‘rse_jx_mouse_RNAseq_nic-smo.Rdata’: (
rse_jxobject) the jx RSE object contains the raw and log-normalized expression data of 1,436,068 mouse exon-exon junctions across the 208 samples from brain and blood of control and nicotine/smoking-exposed pup and adult mice.
All the above datasets contain the sample and feature metadata and additional data of the results obtained in the filtering steps and the DEA.
-
‘de_genes_prenatal_human_brain_smoking.Rdata’:
(
de_genes_prenatal_human_brain_smokingobject) data frame with DE statistics of 18,067 human genes for cigarette smoke exposure in prenatal human cortical tissue. -
‘de_genes_adult_human_brain_smoking.Rdata’:
(
de_genes_adult_human_brain_smokingobject) data frame with DE statistics of 18,067 human genes for cigarette smoke exposure in adult human cortical tissue.
Variables of mice data
Feature information in rowData(RSE) contains the
following variables:
-
retained_after_feature_filtering: Boolean variable that equals TRUE if the feature passed the feature filtering based on expression levels and FALSE if not. Check code for the feature filtering analysis in here. -
DE_in_adult_brain_nicotine: Boolean variable that equals TRUE if the feature is differentially expressed (DE) for nicotine vs vehicle administration in adult brain and FALSE if not. Check code for the DEA in here. -
DE_in_adult_brain_smoking: Boolean variable that equals TRUE if the feature is differentially expressed (DE) for smoking exposure vs control in adult brain and FALSE if not. Check code for the DEA in here. -
DE_in_adult_blood_smoking: Boolean variable that equals TRUE if the feature is differentially expressed (DE) for smoking exposure vs control in adult blood and FALSE if not. Check code for the DEA in here. -
DE_in_pup_brain_nicotine: Boolean variable that equals TRUE if the feature is differentially expressed (DE) for nicotine vs vehicle exposure in pup brain and FALSE if not. Check code for the DEA in here. -
DE_in_pup_brain_smoking: Boolean variable that equals TRUE if the feature is differentially expressed (DE) for smoking exposure vs control in pup brain and FALSE if not. Check code for the DEA in here.
The rest of the variables are outputs of the SPEAQeasy pipeline (Eagles et al., 2021). See here for their description.
Sample information in colData(RSE) contains the
following variables:
- The Quality Control (QC) variables
sum,detected,subsets_Mito_sum,subsets_Mito_detected,subsets_Mito_percent,subsets_Ribo_sum,subsets_Ribo_detected, andsubsets_Ribo_percentare returned byaddPerCellQC()from scuttle. See here for more details. -
retained_after_QC_sample_filtering: Boolean variable that equals TRUE if the sample passed the sample filtering based on QC metrics and FALSE if not. Check code for QC-based sample filtering in here. -
retained_after_manual_sample_filtering: Boolean variable that equals TRUE if the sample passed the manual sample filtering based on PCA plots and FALSE if not. Check code for PCA-based sample filtering in here
The rest of the variables are outputs of SPEAQeasy. See their description here.
Variables of human data
Check Smoking_DLPFC_Devel code for human data generation and limma documentation for the description of differential expression statistics.
Downloading the data with smokingMouse
Using smokingMouse
(Gonzalez-Padilla, Eagles, Cano, Pertea, Jaffe, Maynard, Hancock, Handa,
Martinowich, Maynard, and Collado-Torres, 2024) you can download these
R objects. They are hosted by Bioconductor’s ExperimentHub
(Morgan and Shepherd, 2025) resource. Below you can see how to obtain
these objects.
## Load ExperimentHub for downloading the data
library("ExperimentHub")
#> Loading required package: BiocGenerics
#> Loading required package: generics
#>
#> Attaching package: 'generics'
#> The following objects are masked from 'package:base':
#>
#> as.difftime, as.factor, as.ordered, intersect, is.element, setdiff,
#> setequal, union
#>
#> Attaching package: 'BiocGenerics'
#> The following objects are masked from 'package:stats':
#>
#> IQR, mad, sd, var, xtabs
#> The following objects are masked from 'package:base':
#>
#> anyDuplicated, aperm, append, as.data.frame, basename, cbind,
#> colnames, dirname, do.call, duplicated, eval, evalq, Filter, Find,
#> get, grep, grepl, is.unsorted, lapply, Map, mapply, match, mget,
#> order, paste, pmax, pmax.int, pmin, pmin.int, Position, rank,
#> rbind, Reduce, rownames, sapply, saveRDS, table, tapply, unique,
#> unsplit, which.max, which.min
#> Loading required package: AnnotationHub
#> Loading required package: BiocFileCache
#> Loading required package: dbplyr
#> Registered S3 method overwritten by 'bit64':
#> method from
#> print.bitstring tools
## Connect to ExperimentHub
ehub <- ExperimentHub::ExperimentHub()
## Load the datasets of the package
myfiles <- query(ehub, "smokingMouse")
## Resulting smokingMouse files from our ExperimentHub query
myfiles
#> ExperimentHub with 6 records
#> # snapshotDate(): 2026-03-30
#> # $dataprovider: Lieber Institute for Brain Development (LIBD)
#> # $species: Mus musculus, Homo sapiens
#> # $rdataclass: RangedSummarizedExperiment, GenomicRanges
#> # additional mcols(): taxonomyid, genome, description,
#> # coordinate_1_based, maintainer, rdatadateadded, preparerclass, tags,
#> # rdatapath, sourceurl, sourcetype
#> # retrieve records with, e.g., 'object[["EH8313"]]'
#>
#> title
#> EH8313 | rse_gene_mouse_RNAseq_nic-smo
#> EH8314 | rse_tx_mouse_RNAseq_nic-smo
#> EH8315 | rse_jx_mouse_RNAseq_nic-smo
#> EH8316 | rse_exon_mouse_RNAseq_nic-smo
#> EH8317 | de_genes_prenatal_human_brain_smoking
#> EH8318 | de_genes_adult_human_brain_smoking
## Load SummarizedExperiment which defines the class container for the data
library("SummarizedExperiment")
#> Loading required package: MatrixGenerics
#> Loading required package: matrixStats
#>
#> Attaching package: 'MatrixGenerics'
#> The following objects are masked from 'package:matrixStats':
#>
#> colAlls, colAnyNAs, colAnys, colAvgsPerRowSet, colCollapse,
#> colCounts, colCummaxs, colCummins, colCumprods, colCumsums,
#> colDiffs, colIQRDiffs, colIQRs, colLogSumExps, colMadDiffs,
#> colMads, colMaxs, colMeans2, colMedians, colMins, colOrderStats,
#> colProds, colQuantiles, colRanges, colRanks, colSdDiffs, colSds,
#> colSums2, colTabulates, colVarDiffs, colVars, colWeightedMads,
#> colWeightedMeans, colWeightedMedians, colWeightedSds,
#> colWeightedVars, rowAlls, rowAnyNAs, rowAnys, rowAvgsPerColSet,
#> rowCollapse, rowCounts, rowCummaxs, rowCummins, rowCumprods,
#> rowCumsums, rowDiffs, rowIQRDiffs, rowIQRs, rowLogSumExps,
#> rowMadDiffs, rowMads, rowMaxs, rowMeans2, rowMedians, rowMins,
#> rowOrderStats, rowProds, rowQuantiles, rowRanges, rowRanks,
#> rowSdDiffs, rowSds, rowSums2, rowTabulates, rowVarDiffs, rowVars,
#> rowWeightedMads, rowWeightedMeans, rowWeightedMedians,
#> rowWeightedSds, rowWeightedVars
#> Loading required package: GenomicRanges
#> Loading required package: stats4
#> Loading required package: S4Vectors
#>
#> Attaching package: 'S4Vectors'
#> The following object is masked from 'package:utils':
#>
#> findMatches
#> The following objects are masked from 'package:base':
#>
#> expand.grid, I, unname
#> Loading required package: IRanges
#> Loading required package: Seqinfo
#> Loading required package: Biobase
#> Welcome to Bioconductor
#>
#> Vignettes contain introductory material; view with
#> 'browseVignettes()'. To cite Bioconductor, see
#> 'citation("Biobase")', and for packages 'citation("pkgname")'.
#>
#> Attaching package: 'Biobase'
#> The following object is masked from 'package:MatrixGenerics':
#>
#> rowMedians
#> The following objects are masked from 'package:matrixStats':
#>
#> anyMissing, rowMedians
#> The following object is masked from 'package:ExperimentHub':
#>
#> cache
#> The following object is masked from 'package:AnnotationHub':
#>
#> cache
######################
# Mouse data
######################
myfiles["EH8313"]
#> ExperimentHub with 1 record
#> # snapshotDate(): 2026-03-30
#> # names(): EH8313
#> # package(): smokingMouse
#> # $dataprovider: Lieber Institute for Brain Development (LIBD)
#> # $species: Mus musculus
#> # $rdataclass: RangedSummarizedExperiment
#> # $rdatadateadded: 2023-07-21
#> # $title: rse_gene_mouse_RNAseq_nic-smo
#> # $description: RangedSummarizedExperiment of bulk RNA-seq data from mouse b...
#> # $taxonomyid: 10090
#> # $genome: GRCm38
#> # $sourcetype: GTF
#> # $sourceurl: https://bioconductor.org/packages/smokingMouse
#> # $sourcesize: NA
#> # $tags: c("ExpressionData", "LIBD", "mouse", "Mus_musculus_Data",
#> # "nicotine", "RNASeqData", "smoking")
#> # retrieve record with 'object[["EH8313"]]'
## Download the mouse gene data
# EH8313 | rse_gene_mouse_RNAseq_nic-smo
rse_gene <- myfiles[["EH8313"]]
#> see ?smokingMouse and browseVignettes('smokingMouse') for documentation
#> loading from cache
## This is a RangedSummarizedExperiment object
rse_gene
#> class: RangedSummarizedExperiment
#> dim: 55401 208
#> metadata(1): Obtained_from
#> assays(2): counts logcounts
#> rownames(55401): ENSMUSG00000102693.1 ENSMUSG00000064842.1 ...
#> ENSMUSG00000064371.1 ENSMUSG00000064372.1
#> rowData names(13): Length gencodeID ... DE_in_pup_brain_nicotine
#> DE_in_pup_brain_smoking
#> colnames: NULL
#> colData names(71): SAMPLE_ID FQCbasicStats ...
#> retained_after_QC_sample_filtering
#> retained_after_manual_sample_filtering
## Optionally check the memory size
# lobstr::obj_size(rse_gene)
# 159.68 MB
## Check sample info
head(colData(rse_gene), 3)
#> DataFrame with 3 rows and 71 columns
#> SAMPLE_ID FQCbasicStats perBaseQual perTileQual perSeqQual perBaseContent
#> <character> <character> <character> <character> <character> <character>
#> 1 Sample_2914 PASS PASS PASS PASS FAIL/WARN
#> 2 Sample_4042 PASS PASS PASS PASS FAIL/WARN
#> 3 Sample_4043 PASS PASS PASS PASS FAIL/WARN
#> GCcontent Ncontent SeqLengthDist SeqDuplication OverrepSeqs
#> <character> <character> <character> <character> <character>
#> 1 WARN PASS WARN FAIL PASS
#> 2 WARN PASS WARN FAIL PASS
#> 3 WARN PASS WARN FAIL PASS
#> AdapterContent KmerContent SeqLength_R1 percentGC_R1 phred15-19_R1
#> <character> <character> <character> <character> <character>
#> 1 PASS NA 75-151 49 37.0
#> 2 PASS NA 75-151 48 37.0
#> 3 PASS NA 75-151 49 37.0
#> phred65-69_R1 phred115-119_R1 phred150-151_R1 phredGT30_R1 phredGT35_R1
#> <character> <character> <character> <numeric> <numeric>
#> 1 37.0 37.0 37.0 NA NA
#> 2 37.0 37.0 37.0 NA NA
#> 3 37.0 37.0 37.0 NA NA
#> Adapter65-69_R1 Adapter100-104_R1 Adapter140_R1 SeqLength_R2 percentGC_R2
#> <numeric> <numeric> <numeric> <character> <character>
#> 1 0.000108294 0.000260890 0.00174971 75-151 49
#> 2 0.000210067 0.000352780 0.00154716 75-151 48
#> 3 0.000134434 0.000284284 0.00191475 75-151 49
#> phred15-19_R2 phred65-69_R2 phred115-119_R2 phred150-151_R2 phredGT30_R2
#> <character> <character> <character> <character> <numeric>
#> 1 37.0 37.0 37.0 37.0 NA
#> 2 37.0 37.0 37.0 37.0 NA
#> 3 37.0 37.0 37.0 37.0 NA
#> phredGT35_R2 Adapter65-69_R2 Adapter100-104_R2 Adapter140_R2 ERCCsumLogErr
#> <numeric> <numeric> <numeric> <numeric> <numeric>
#> 1 NA 0.000276104 0.000486875 0.00132235 -58.2056
#> 2 NA 0.000326771 0.000574851 0.00137044 -81.6359
#> 3 NA 0.000330534 0.000521084 0.00153550 -99.5348
#> bamFile trimmed numReads numMapped numUnmapped
#> <character> <logical> <numeric> <numeric> <numeric>
#> 1 Sample_2914_sorted.bam FALSE 89386472 87621022 1765450
#> 2 Sample_4042_sorted.bam FALSE 59980794 58967812 1012982
#> 3 Sample_4043_sorted.bam FALSE 64864732 63961359 903373
#> overallMapRate concordMapRate totalMapped mitoMapped mitoRate
#> <numeric> <numeric> <numeric> <numeric> <numeric>
#> 1 0.9802 0.9748 87143087 10039739 0.103308
#> 2 0.9831 0.9709 58215746 7453208 0.113497
#> 3 0.9861 0.9811 62983384 8307331 0.116528
#> totalAssignedGene rRNA_rate Tissue Age Sex Expt
#> <numeric> <numeric> <character> <character> <character> <character>
#> 1 0.761378 0.00396315 Brain Adult F Nicotine
#> 2 0.754444 0.00301119 Brain Adult F Nicotine
#> 3 0.757560 0.00288706 Brain Adult F Nicotine
#> Group Pregnant plate location concentration medium
#> <character> <character> <character> <character> <character> <character>
#> 1 Experimental 0 Plate2 C12 165.9 Water
#> 2 Control 0 Plate1 B4 122.6 Water
#> 3 Control 0 Plate2 C9 128.5 Water
#> date Pregnancy flowcell sum detected subsets_Mito_sum
#> <character> <character> <character> <numeric> <numeric> <numeric>
#> 1 NA No HKCMHDSXX 37119948 24435 2649559
#> 2 NA No HKCG7DSXX 24904754 23656 1913803
#> 3 NA No HKCMHDSXX 27083602 23903 2180712
#> subsets_Mito_detected subsets_Mito_percent subsets_Ribo_sum
#> <numeric> <numeric> <numeric>
#> 1 26 7.13783 486678
#> 2 31 7.68449 319445
#> 3 31 8.05178 338277
#> subsets_Ribo_detected subsets_Ribo_percent retained_after_QC_sample_filtering
#> <numeric> <numeric> <logical>
#> 1 11 1.31110 TRUE
#> 2 13 1.28267 TRUE
#> 3 14 1.24901 TRUE
#> retained_after_manual_sample_filtering
#> <logical>
#> 1 TRUE
#> 2 TRUE
#> 3 TRUE
## Check gene info
head(rowData(rse_gene), 3)
#> DataFrame with 3 rows and 13 columns
#> Length gencodeID ensemblID
#> <integer> <character> <character>
#> ENSMUSG00000102693.1 1070 ENSMUSG00000102693.1 ENSMUSG00000102693
#> ENSMUSG00000064842.1 110 ENSMUSG00000064842.1 ENSMUSG00000064842
#> ENSMUSG00000051951.5 6094 ENSMUSG00000051951.5 ENSMUSG00000051951
#> gene_type EntrezID Symbol meanExprs
#> <character> <character> <character> <numeric>
#> ENSMUSG00000102693.1 TEC 71042 MGI:MGI:1918292 0.00000
#> ENSMUSG00000064842.1 snRNA NA NA 0.00000
#> ENSMUSG00000051951.5 protein_coding 497097 MGI:MGI:3528744 7.94438
#> retained_after_feature_filtering DE_in_adult_blood_smoking
#> <logical> <logical>
#> ENSMUSG00000102693.1 FALSE FALSE
#> ENSMUSG00000064842.1 FALSE FALSE
#> ENSMUSG00000051951.5 TRUE FALSE
#> DE_in_adult_brain_nicotine DE_in_adult_brain_smoking
#> <logical> <logical>
#> ENSMUSG00000102693.1 FALSE FALSE
#> ENSMUSG00000064842.1 FALSE FALSE
#> ENSMUSG00000051951.5 FALSE FALSE
#> DE_in_pup_brain_nicotine DE_in_pup_brain_smoking
#> <logical> <logical>
#> ENSMUSG00000102693.1 FALSE FALSE
#> ENSMUSG00000064842.1 FALSE FALSE
#> ENSMUSG00000051951.5 FALSE FALSE
## Access the original counts
class(assays(rse_gene)$counts)
#> [1] "matrix" "array"
dim(assays(rse_gene)$counts)
#> [1] 55401 208
assays(rse_gene)$counts[1:3, 1:3]
#> [,1] [,2] [,3]
#> ENSMUSG00000102693.1 0 0 0
#> ENSMUSG00000064842.1 0 0 0
#> ENSMUSG00000051951.5 811 710 812
## Access the log-normalized counts
class(assays(rse_gene)$logcounts)
#> [1] "matrix" "array"
assays(rse_gene)$logcounts[1:3, 1:3]
#> [,1] [,2] [,3]
#> ENSMUSG00000102693.1 -5.985331 -5.985331 -5.985331
#> ENSMUSG00000064842.1 -5.985331 -5.985331 -5.985331
#> ENSMUSG00000051951.5 4.509114 4.865612 4.944597
######################
# Human data
######################
myfiles["EH8318"]
#> ExperimentHub with 1 record
#> # snapshotDate(): 2026-03-30
#> # names(): EH8318
#> # package(): smokingMouse
#> # $dataprovider: Lieber Institute for Brain Development (LIBD)
#> # $species: Homo sapiens
#> # $rdataclass: GenomicRanges
#> # $rdatadateadded: 2023-07-21
#> # $title: de_genes_adult_human_brain_smoking
#> # $description: GRanges with the information of the differentialy expressed ...
#> # $taxonomyid: 9606
#> # $genome: GRCh37
#> # $sourcetype: GTF
#> # $sourceurl: https://bioconductor.org/packages/smokingMouse
#> # $sourcesize: NA
#> # $tags: c("ExpressionData", "LIBD", "mouse", "Mus_musculus_Data",
#> # "nicotine", "RNASeqData", "smoking")
#> # retrieve record with 'object[["EH8318"]]'
## Download the human gene data
# EH8318 | de_genes_adult_human_brain_smoking
de_genes_prenatal_human_brain_smoking <- myfiles[["EH8318"]]
#> see ?smokingMouse and browseVignettes('smokingMouse') for documentation
#> loading from cache
## This is a GRanges object
class(de_genes_prenatal_human_brain_smoking)
#> [1] "GRanges"
#> attr(,"package")
#> [1] "GenomicRanges"
de_genes_prenatal_human_brain_smoking
#> GRanges object with 18067 ranges and 9 metadata columns:
#> seqnames ranges strand | Length Symbol
#> <Rle> <IRanges> <Rle> | <integer> <character>
#> ENSG00000019169 chr2 119699742-119752236 + | 2079 MARCO
#> ENSG00000260400 chr10 70458257-70460551 + | 2295
#> ENSG00000011201 chrX 8496915-8700227 - | 7131 KAL1
#> ENSG00000068438 chrX 48334541-48344752 + | 2740 FTSJ1
#> ENSG00000151229 chr12 40148823-40499891 - | 10027 SLC2A13
#> ... ... ... ... . ... ...
#> ENSG00000141556 chr17 80709940-80900724 + | 10472 TBCD
#> ENSG00000125804 chr20 26035291-26073683 + | 6332 FAM182A
#> ENSG00000228998 chr15 90818266-90820841 + | 2576
#> ENSG00000149636 chr20 35380194-35402221 - | 2739 DSN1
#> ENSG00000122644 chr7 12726481-12730559 + | 3561 ARL4A
#> EntrezID logFC AveExpr t P.Value
#> <integer> <numeric> <numeric> <numeric> <numeric>
#> ENSG00000019169 8685 -1.6032766 -1.80183 -6.14514 4.66989e-09
#> ENSG00000260400 <NA> 0.1515813 1.17142 4.09836 6.18298e-05
#> ENSG00000011201 3730 0.1423143 4.24576 4.09392 6.29277e-05
#> ENSG00000068438 24140 -0.0495086 4.30660 -4.05975 7.20166e-05
#> ENSG00000151229 114134 0.0842742 7.02625 4.00115 9.05839e-05
#> ... ... ... ... ... ...
#> ENSG00000141556 101929597 -9.07984e-06 6.16583 -4.49543e-04 0.999642
#> ENSG00000125804 728882 -2.02864e-05 2.75543 -2.93079e-04 0.999766
#> ENSG00000228998 <NA> 1.05697e-05 4.14580 2.50417e-04 0.999800
#> ENSG00000149636 79980 2.46976e-06 2.96401 1.02959e-04 0.999918
#> ENSG00000122644 10124 1.62178e-06 4.51605 4.83868e-05 0.999961
#> adj.P.Val B
#> <numeric> <numeric>
#> ENSG00000019169 0.000084371 3.65960
#> ENSG00000260400 0.325280946 1.21448
#> ENSG00000011201 0.325280946 1.55562
#> ENSG00000068438 0.325280946 1.43237
#> ENSG00000151229 0.327315930 1.15523
#> ... ... ...
#> ENSG00000141556 0.999863 -6.30298
#> ENSG00000125804 0.999911 -5.80161
#> ENSG00000228998 0.999911 -6.07834
#> ENSG00000149636 0.999961 -5.84217
#> ENSG00000122644 0.999961 -6.13228
#> -------
#> seqinfo: 25 sequences from an unspecified genome; no seqlengths
## Optionally check the memory size
# lobstr::obj_size(de_genes_prenatal_human_brain_smoking)
# 3.73 MB
## Access data of human genes as normally do with other GenomicRanges::GRanges()
## objects or re-cast it as a data.frame
de_genes_df <- as.data.frame(de_genes_prenatal_human_brain_smoking)
head(de_genes_df)
#> seqnames start end width strand Length Symbol
#> ENSG00000019169 chr2 119699742 119752236 52495 + 2079 MARCO
#> ENSG00000260400 chr10 70458257 70460551 2295 + 2295
#> ENSG00000011201 chrX 8496915 8700227 203313 - 7131 KAL1
#> ENSG00000068438 chrX 48334541 48344752 10212 + 2740 FTSJ1
#> ENSG00000151229 chr12 40148823 40499891 351069 - 10027 SLC2A13
#> ENSG00000136319 chr14 20724717 20774153 49437 - 9705 TTC5
#> EntrezID logFC AveExpr t P.Value
#> ENSG00000019169 8685 -1.60327659 -1.801830 -6.145143 4.669893e-09
#> ENSG00000260400 NA 0.15158127 1.171418 4.098360 6.182984e-05
#> ENSG00000011201 3730 0.14231431 4.245759 4.093917 6.292772e-05
#> ENSG00000068438 24140 -0.04950860 4.306597 -4.059746 7.201659e-05
#> ENSG00000151229 114134 0.08427416 7.026246 4.001149 9.058392e-05
#> ENSG00000136319 91875 -0.09445164 3.683599 -3.931427 1.186235e-04
#> adj.P.Val B
#> ENSG00000019169 8.437096e-05 3.659604
#> ENSG00000260400 3.252809e-01 1.214477
#> ENSG00000011201 3.252809e-01 1.555617
#> ENSG00000068438 3.252809e-01 1.432373
#> ENSG00000151229 3.273159e-01 1.155227
#> ENSG00000136319 3.497224e-01 1.000380Reproducibility
The smokingMouse package (Gonzalez-Padilla, Eagles, Cano et al., 2024) and the study analyses were made possible thanks to:
-
R(R Core Team, 2026) - AnnotationHubData (Bioconductor Package Maintainer, 2025)
- BiocStyle (Oleś, 2025)
- ExperimentHub (Morgan and Shepherd, 2025)
- ExperimentHubData (Maintainer, 2025)
- knitr (Xie, 2025)
- RefManageR (McLean, 2017)
- rmarkdown (Allaire, Xie, Dervieux, McPherson, Luraschi, Ushey, Atkins, Wickham, Cheng, Chang, and Iannone, 2026)
- sessioninfo (Wickham, Chang, Flight, Müller, and Hester, 2025)
- testthat (Wickham, 2011)
This package was developed using biocthis.
Date the vignette was generated.
#> [1] "2026-03-31 17:53:31 UTC"Wallclock time spent generating the vignette.
#> Time difference of 11.731 secsR session information.
#> ─ Session info ───────────────────────────────────────────────────────────────────────────────────────────────────────
#> setting value
#> version R Under development (unstable) (2026-03-28 r89738)
#> os Ubuntu 24.04.4 LTS
#> system x86_64, linux-gnu
#> ui X11
#> language en
#> collate en_US.UTF-8
#> ctype en_US.UTF-8
#> tz UTC
#> date 2026-03-31
#> pandoc 3.9.0.2 @ /usr/bin/ (via rmarkdown)
#> quarto 1.8.25 @ /usr/local/bin/quarto
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> abind 1.4-8 2024-09-12 [1] CRAN (R 4.6.0)
#> AnnotationDbi 1.73.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> AnnotationHub * 4.1.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> backports 1.5.0 2024-05-23 [1] CRAN (R 4.6.0)
#> bibtex 0.5.2 2026-02-03 [1] CRAN (R 4.6.0)
#> Biobase * 2.71.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> BiocFileCache * 3.1.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> BiocGenerics * 0.57.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> BiocManager 1.30.27 2025-11-14 [2] CRAN (R 4.7.0)
#> BiocStyle * 2.39.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> BiocVersion 3.23.1 2025-10-30 [2] Bioconductor 3.23 (R 4.7.0)
#> Biostrings 2.79.5 2026-03-06 [1] Bioconductor 3.23 (R 4.7.0)
#> bit 4.6.0 2025-03-06 [1] CRAN (R 4.6.0)
#> bit64 4.6.0-1 2025-01-16 [1] CRAN (R 4.6.0)
#> blob 1.3.0 2026-01-14 [1] CRAN (R 4.6.0)
#> bookdown 0.46 2025-12-05 [1] CRAN (R 4.6.0)
#> bslib 0.10.0 2026-01-26 [2] CRAN (R 4.7.0)
#> cachem 1.1.0 2024-05-16 [2] CRAN (R 4.7.0)
#> cli 3.6.5 2025-04-23 [2] CRAN (R 4.7.0)
#> crayon 1.5.3 2024-06-20 [2] CRAN (R 4.7.0)
#> curl 7.0.0 2025-08-19 [2] CRAN (R 4.7.0)
#> DBI 1.3.0 2026-02-25 [1] CRAN (R 4.7.0)
#> dbplyr * 2.5.2 2026-02-13 [1] CRAN (R 4.7.0)
#> DelayedArray 0.37.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> desc 1.4.3 2023-12-10 [2] CRAN (R 4.7.0)
#> digest 0.6.39 2025-11-19 [2] CRAN (R 4.7.0)
#> dplyr 1.2.0 2026-02-03 [1] CRAN (R 4.6.0)
#> evaluate 1.0.5 2025-08-27 [2] CRAN (R 4.7.0)
#> ExperimentHub * 3.1.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> fastmap 1.2.0 2024-05-15 [2] CRAN (R 4.7.0)
#> filelock 1.0.3 2023-12-11 [1] CRAN (R 4.6.0)
#> fs 2.0.1 2026-03-24 [2] CRAN (R 4.7.0)
#> generics * 0.1.4 2025-05-09 [1] CRAN (R 4.6.0)
#> GenomicRanges * 1.63.1 2025-12-08 [1] Bioconductor 3.23 (R 4.6.0)
#> glue 1.8.0 2024-09-30 [2] CRAN (R 4.7.0)
#> htmltools 0.5.9 2025-12-04 [2] CRAN (R 4.7.0)
#> htmlwidgets 1.6.4 2023-12-06 [2] CRAN (R 4.7.0)
#> httr 1.4.8 2026-02-13 [1] CRAN (R 4.7.0)
#> httr2 1.2.2 2025-12-08 [2] CRAN (R 4.7.0)
#> IRanges * 2.45.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> jquerylib 0.1.4 2021-04-26 [2] CRAN (R 4.7.0)
#> jsonlite 2.0.0 2025-03-27 [2] CRAN (R 4.7.0)
#> KEGGREST 1.51.1 2025-11-17 [1] Bioconductor 3.23 (R 4.6.0)
#> knitr 1.51 2025-12-20 [2] CRAN (R 4.7.0)
#> lattice 0.22-9 2026-02-09 [3] CRAN (R 4.7.0)
#> lifecycle 1.0.5 2026-01-08 [2] CRAN (R 4.7.0)
#> lubridate 1.9.5 2026-02-04 [1] CRAN (R 4.6.0)
#> magrittr 2.0.4 2025-09-12 [2] CRAN (R 4.7.0)
#> Matrix 1.7-5 2026-03-21 [3] CRAN (R 4.7.0)
#> MatrixGenerics * 1.23.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> matrixStats * 1.5.0 2025-01-07 [1] CRAN (R 4.6.0)
#> memoise 2.0.1 2021-11-26 [2] CRAN (R 4.7.0)
#> otel 0.2.0 2025-08-29 [2] CRAN (R 4.7.0)
#> pillar 1.11.1 2025-09-17 [2] CRAN (R 4.7.0)
#> pkgconfig 2.0.3 2019-09-22 [2] CRAN (R 4.7.0)
#> pkgdown 2.2.0.9000 2026-03-31 [1] Github (r-lib/pkgdown@a6abe43)
#> plyr 1.8.9 2023-10-02 [1] CRAN (R 4.6.0)
#> png 0.1-9 2026-03-15 [1] CRAN (R 4.7.0)
#> purrr 1.2.1 2026-01-09 [2] CRAN (R 4.7.0)
#> R6 2.6.1 2025-02-15 [2] CRAN (R 4.7.0)
#> ragg 1.5.2 2026-03-23 [2] CRAN (R 4.7.0)
#> rappdirs 0.3.4 2026-01-17 [2] CRAN (R 4.7.0)
#> Rcpp 1.1.1 2026-01-10 [2] CRAN (R 4.7.0)
#> RefManageR * 1.4.0 2022-09-30 [1] CRAN (R 4.6.0)
#> rlang 1.1.7 2026-01-09 [2] CRAN (R 4.7.0)
#> rmarkdown 2.31 2026-03-26 [2] CRAN (R 4.7.0)
#> RSQLite 2.4.6 2026-02-06 [1] CRAN (R 4.6.0)
#> S4Arrays 1.11.1 2025-11-25 [1] Bioconductor 3.23 (R 4.6.0)
#> S4Vectors * 0.49.0 2025-10-30 [1] Bioconductor 3.23 (R 4.6.0)
#> sass 0.4.10 2025-04-11 [2] CRAN (R 4.7.0)
#> Seqinfo * 1.1.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> sessioninfo * 1.2.3 2025-02-05 [2] CRAN (R 4.7.0)
#> smokingMouse * 1.9.0 2026-03-31 [1] Bioconductor
#> SparseArray 1.11.12 2026-03-30 [1] Bioconductor 3.23 (R 4.7.0)
#> stringi 1.8.7 2025-03-27 [2] CRAN (R 4.7.0)
#> stringr 1.6.0 2025-11-04 [2] CRAN (R 4.7.0)
#> SummarizedExperiment * 1.41.1 2026-02-06 [1] Bioconductor 3.23 (R 4.6.0)
#> systemfonts 1.3.2 2026-03-05 [2] CRAN (R 4.7.0)
#> textshaping 1.0.5 2026-03-06 [2] CRAN (R 4.7.0)
#> tibble 3.3.1 2026-01-11 [2] CRAN (R 4.7.0)
#> tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.6.0)
#> timechange 0.4.0 2026-01-29 [1] CRAN (R 4.6.0)
#> vctrs 0.7.2 2026-03-21 [2] CRAN (R 4.7.0)
#> withr 3.0.2 2024-10-28 [2] CRAN (R 4.7.0)
#> xfun 0.57 2026-03-20 [2] CRAN (R 4.7.0)
#> xml2 1.5.2 2026-01-17 [2] CRAN (R 4.7.0)
#> XVector 0.51.0 2025-10-31 [1] Bioconductor 3.23 (R 4.6.0)
#> yaml 2.3.12 2025-12-10 [2] CRAN (R 4.7.0)
#>
#> [1] /__w/_temp/Library
#> [2] /usr/local/lib/R/site-library
#> [3] /usr/local/lib/R/library
#> * ── Packages attached to the search path.
#>
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────Bibliography
This vignette was generated using BiocStyle (Oleś, 2025) with knitr (Xie, 2025) and rmarkdown (Allaire, Xie, Dervieux et al., 2026) running behind the scenes.
Citations made with RefManageR (McLean, 2017).
[1] J. Allaire, Y. Xie, C. Dervieux, et al. rmarkdown: Dynamic Documents for R. R package version 2.31. 2026. URL: https://github.com/rstudio/rmarkdown.
[2] Bioconductor Package Maintainer. AnnotationHubData: Transform public data resources into Bioconductor Data Structures. R package version 1.41.0. 2025. DOI: 10.18129/B9.bioc.AnnotationHubData. URL: https://bioconductor.org/packages/AnnotationHubData.
[3] D. Gonzalez-Padilla, N. J. Eagles, M. Cano, et al. “Molecular impact of nicotine and smoking exposure on the developing and adult mouse brain”. In: bioRxiv (2024). DOI: 10.1101/2024.11.05.622149. URL: https://doi.org/10.1101/2024.11.05.622149.
[4] B. Maintainer. ExperimentHubData: Add resources to ExperimentHub. R package version 1.37.0. 2025. DOI: 10.18129/B9.bioc.ExperimentHubData. URL: https://bioconductor.org/packages/ExperimentHubData.
[5] M. W. McLean. “RefManageR: Import and Manage BibTeX and BibLaTeX References in R”. In: The Journal of Open Source Software (2017). DOI: 10.21105/joss.00338.
[6] M. Morgan and L. Shepherd. ExperimentHub: Client to access ExperimentHub resources. R package version 3.1.0. 2025. DOI: 10.18129/B9.bioc.ExperimentHub. URL: https://bioconductor.org/packages/ExperimentHub.
[7] A. Oleś. BiocStyle: Standard styles for vignettes and other Bioconductor documents. R package version 2.39.0. 2025. DOI: 10.18129/B9.bioc.BiocStyle. URL: https://bioconductor.org/packages/BiocStyle.
[8] R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing (ROR: <https://ror.org/05qewa988>;). Vienna, Austria, 2026. DOI: 10.32614/R.manuals. URL: https://www.R-project.org/.
[9] H. Wickham. “testthat: Get Started with Testing”. In: The R Journal 3 (2011), pp. 5–10. URL: https://journal.r-project.org/articles/RJ-2011-002/.
[10] H. Wickham, W. Chang, R. Flight, et al. sessioninfo: R Session Information. R package version 1.2.3. 2025. DOI: 10.32614/CRAN.package.sessioninfo. URL: https://CRAN.R-project.org/package=sessioninfo.
[11] Y. Xie. knitr: A General-Purpose Package for Dynamic Report Generation in R. R package version 1.51. 2025. URL: https://yihui.org/knitr/.