This function visualizes the gene expression stored in assays(spe) or any
continuous variable stored in colData(spe) for a set of samples at the
spot-level using (by default) the histology information on the background.
To visualize clusters (or any discrete variable) use vis_grid_clus().
vis_grid_gene(
spe,
geneid = rowData(spe)$gene_search[1],
pdf_file,
assayname = "logcounts",
minCount = 0,
return_plots = FALSE,
spatial = TRUE,
viridis = TRUE,
height = 24,
width = 36,
image_id = "lowres",
alpha = NA,
cont_colors = if (viridis) viridisLite::viridis(21) else c("aquamarine4",
"springgreen", "goldenrod", "red"),
sample_order = unique(spe$sample_id),
point_size = 2,
auto_crop = TRUE,
na_color = "#CCCCCC40",
is_stitched = FALSE,
cap_percentile = 1,
...
)Arguments
- spe
A SpatialExperiment-class object. See
fetch_data()for how to download some example objects orread10xVisiumWrapper()to read inspaceranger --countoutput files and build your ownspeobject.- geneid
A
character()specifying the gene ID(s) stored inrowData(spe)$gene_searchor a continuous variable(s) stored incolData(spe)to visualize. For each ID, ifrowData(spe)$gene_searchis missing, thenrownames(spe)is used to search for the gene ID. When a vector of length > 1 is supplied, the continuous variables are combined according tomulti_gene_method, producing a single value for each spot.- pdf_file
A
character(1)specifying the path for the resulting PDF.- assayname
The name of the
assays(spe)to use for extracting the gene expression data. Defaults tologcounts.- minCount
A
numeric(1)specifying the minimum gene expression (or value in the continuous variable) to visualize. Values at or below this threshold will be set toNA. Defaults to0.- return_plots
A
logical(1)indicating whether to print the plots to a PDF or to return the list of plots that you can then print using plot_grid.- spatial
A
logical(1)indicating whether to include the histology layer fromgeom_spatial(). If you plan to use ggplotly() then it's best to set this toFALSE.- viridis
A
logical(1)whether to use the color-blind friendly palette from viridis or the color palette used in the paper that was chosen for contrast when visualizing the data on top of the histology image. One issue is being able to differentiate low values from NA ones due to the purple-ish histology information that is dependent on cell density.- height
A
numeric(1)passed to pdf.- width
A
numeric(1)passed to pdf.- image_id
A
character(1)with the name of the image ID you want to use in the background.- alpha
A
numeric(1)in the[0, 1]range that specifies the transparency level of the data on the spots.- cont_colors
A
character()vector of colors that supersedes theviridisargument.- sample_order
A
character()with the names of the samples to use and their order.- point_size
A
numeric(1)specifying the size of the points. Defaults to1.25. Some colors look better if you use2for instance.- auto_crop
A
logical(1)indicating whether to automatically crop the image / plotting area, which is useful if the Visium capture area is not centered on the image and if the image is not a square.- na_color
A
character(1)specifying a color for the NA values. If you setalpha = NAthen it's best to setna_colorto a color that has alpha blending already, which will make non-NA values pop up more and the NA values will show with a lighter color. This behavior is lost whenalphais set to a non-NAvalue.- is_stitched
A
logical(1)vector: IfTRUE, expects a SpatialExperiment-class built withvisiumStitched::build_spe(). http://research.libd.org/visiumStitched/reference/build_spe.html; in particular, expects a logical colData columnexclude_overlappingspecifying which spots to exclude from the plot. Setsauto_crop = FALSE.- cap_percentile
A
numeric(1)in (0, 1] determining the maximum percentile (as a proportion) at which to cap expression. For example, a value of 0.95 sets the top 5% of expression values to the 95th percentile value. This can help make the color scale more dynamic in the presence of high outliers. Defaults to1, which effectively performs no capping.- ...
Passed to paste0() for making the title of the plot following the
sampleid.
Value
A list of ggplot2 objects.
Details
This function prepares the data and then loops through
vis_gene() for computing the list of ggplot2
objects.
See also
Other Spatial gene visualization functions:
vis_gene(),
vis_gene_p()
Examples
if (enough_ram()) {
## Obtain the necessary data
if (!exists("spe")) spe <- fetch_data("spe")
## Subset to two samples of interest and obtain the plot list
p_list <-
vis_grid_gene(
spe[, spe$sample_id %in% c("151673", "151674")],
spatial = FALSE,
return_plots = TRUE
)
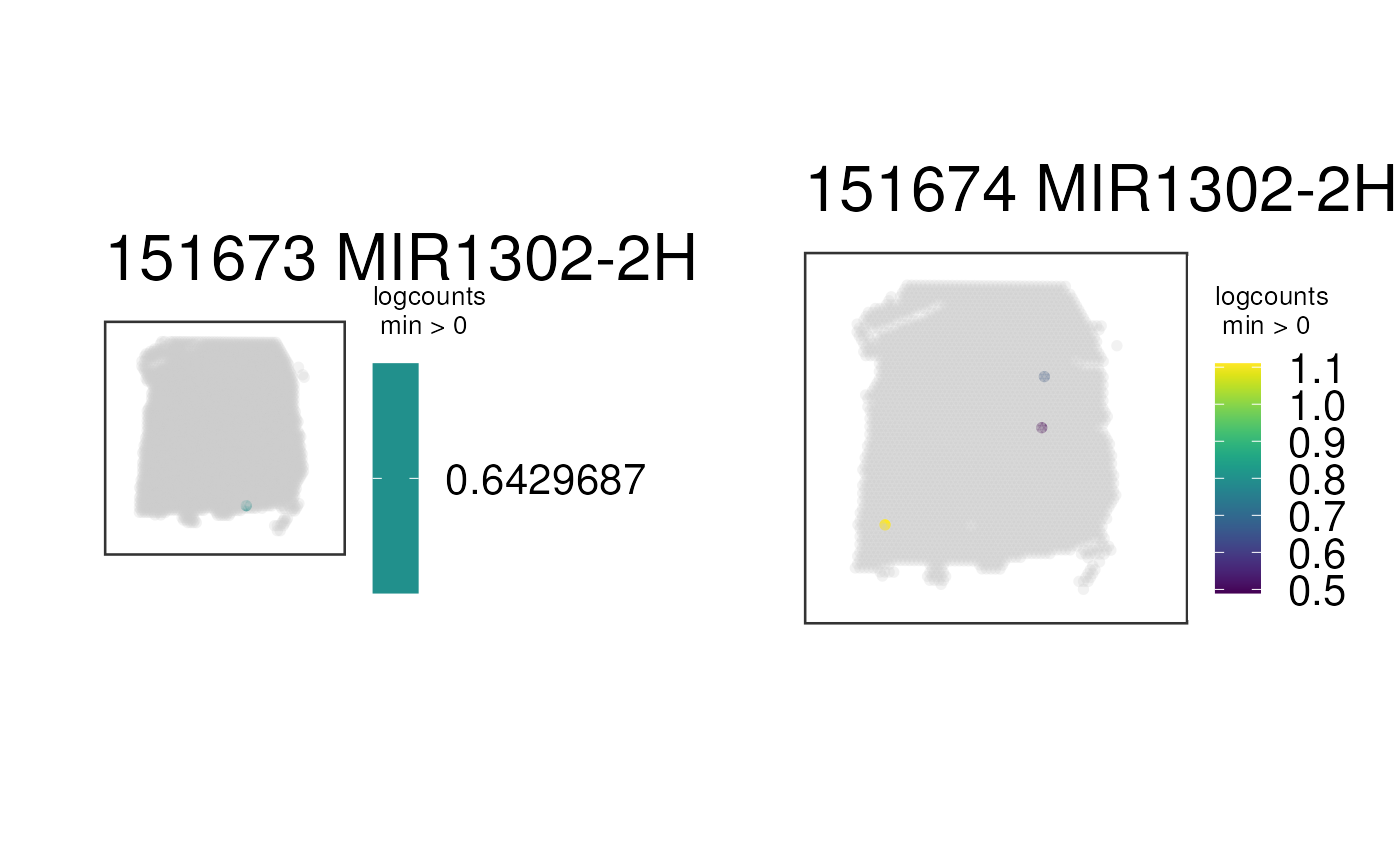
## Visualize the spatial adjacent replicates for position = 0 micro meters
## for subject 3
cowplot::plot_grid(plotlist = p_list, ncol = 2)
}
#> 2026-03-27 00:08:31.133522 loading file /github/home/.cache/R/BiocFileCache/1a6c5ad86da_Human_DLPFC_Visium_processedData_sce_scran_spatialLIBD.Rdata%3Fdl%3D1