This function visualizes the clusters for a set of samples at the spot-level
using (by default) the histology information on the background. To visualize
gene-level (or any continuous variable) use vis_grid_gene().
vis_grid_clus(
spe,
clustervar,
pdf_file,
sort_clust = TRUE,
colors = NULL,
return_plots = FALSE,
spatial = TRUE,
height = 24,
width = 36,
image_id = "lowres",
alpha = NA,
sample_order = unique(spe$sample_id),
point_size = 2,
auto_crop = TRUE,
na_color = "#CCCCCC40",
is_stitched = FALSE,
guide_point_size = point_size,
...
)Arguments
- spe
A SpatialExperiment-class object. See
fetch_data()for how to download some example objects orread10xVisiumWrapper()to read inspaceranger --countoutput files and build your ownspeobject.- clustervar
A
character(1)with the name of thecolData(spe)column that has the cluster values.- pdf_file
A
character(1)specifying the path for the resulting PDF.- sort_clust
A
logical(1)indicating whether you want to sort the clusters by frequency usingsort_clusters().- colors
A vector of colors to use for visualizing the clusters from
clustervar. If the vector has names, then those should match the values ofclustervar.- return_plots
A
logical(1)indicating whether to print the plots to a PDF or to return the list of plots that you can then print using plot_grid.- spatial
A
logical(1)indicating whether to include the histology layer fromgeom_spatial(). If you plan to use ggplotly() then it's best to set this toFALSE.- height
A
numeric(1)passed to pdf.- width
A
numeric(1)passed to pdf.- image_id
A
character(1)with the name of the image ID you want to use in the background.- alpha
A
numeric(1)in the[0, 1]range that specifies the transparency level of the data on the spots.- sample_order
A
character()with the names of the samples to use and their order.- point_size
A
numeric(1)specifying the size of the points. Defaults to1.25. Some colors look better if you use2for instance.- auto_crop
A
logical(1)indicating whether to automatically crop the image / plotting area, which is useful if the Visium capture area is not centered on the image and if the image is not a square.- na_color
A
character(1)specifying a color for the NA values. If you setalpha = NAthen it's best to setna_colorto a color that has alpha blending already, which will make non-NA values pop up more and the NA values will show with a lighter color. This behavior is lost whenalphais set to a non-NAvalue.- is_stitched
A
logical(1)vector: IfTRUE, expects a SpatialExperiment-class built withvisiumStitched::build_spe(). http://research.libd.org/visiumStitched/reference/build_spe.html; in particular, expects a logical colData columnexclude_overlappingspecifying which spots to exclude from the plot. Setsauto_crop = FALSE.- guide_point_size
A
numeric(1)specifying the size of the points in guide. Defaults topoint_size. Increase to improve visability.- ...
Passed to paste0() for making the title of the plot following the
sampleid.
Value
A list of ggplot2 objects.
Details
This function prepares the data and then loops through
vis_clus() for computing the list of ggplot2
objects.
See also
Other Spatial cluster visualization functions:
frame_limits(),
vis_clus(),
vis_clus_p(),
vis_image()
Examples
if (enough_ram()) {
## Obtain the necessary data
if (!exists("spe")) spe <- fetch_data("spe")
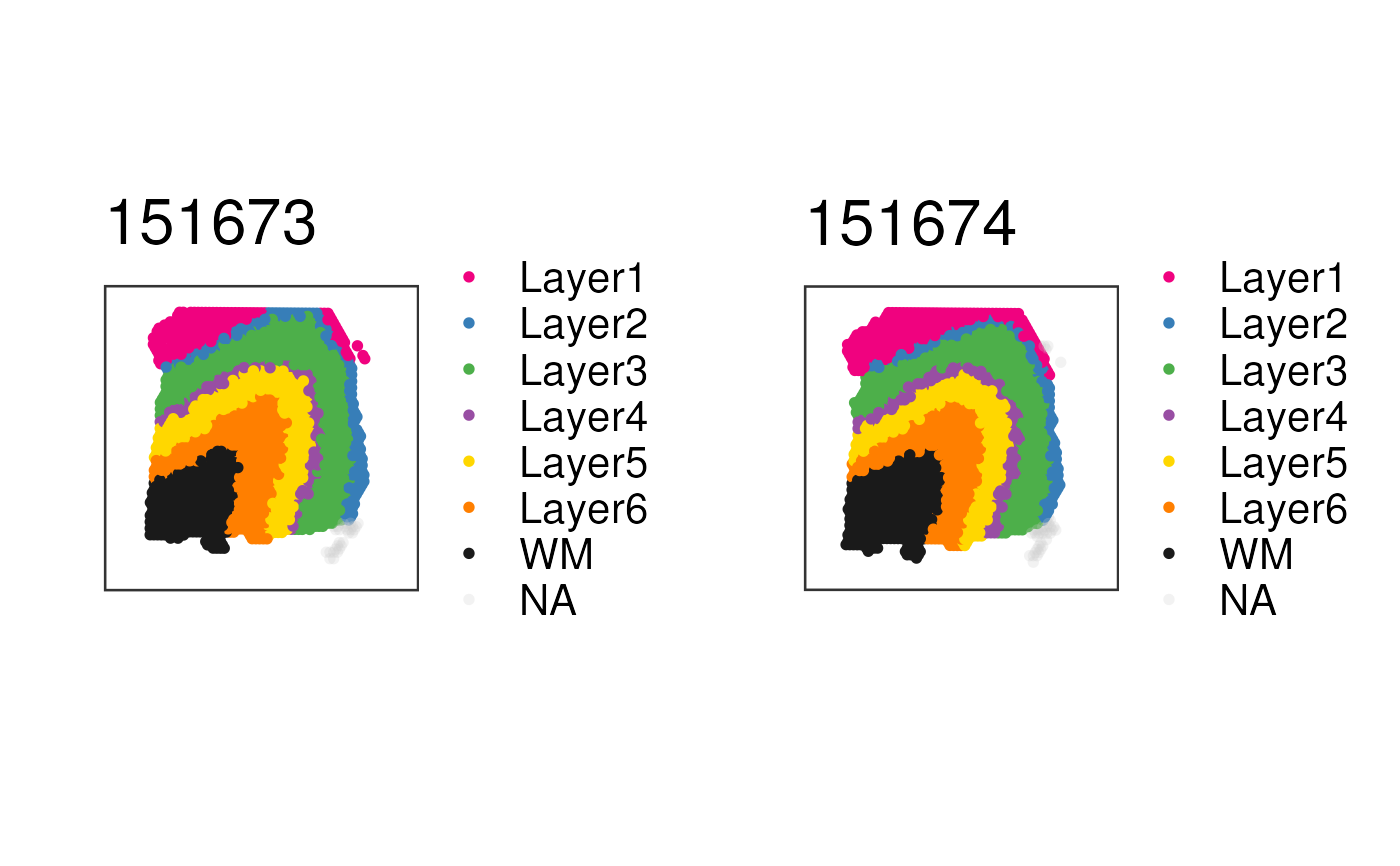
## Subset to two samples of interest and obtain the plot list
p_list <-
vis_grid_clus(
spe[, spe$sample_id %in% c("151673", "151674")],
"layer_guess_reordered",
spatial = FALSE,
return_plots = TRUE,
sort_clust = FALSE,
colors = libd_layer_colors
)
## Visualize the spatial adjacent replicates for position = 0 micro meters
## for subject 3
cowplot::plot_grid(plotlist = p_list, ncol = 2)
}
#> 2026-01-09 17:24:37.150514 loading file /github/home/.cache/R/BiocFileCache/1b5c76453a2f_Human_DLPFC_Visium_processedData_sce_scran_spatialLIBD.Rdata%3Fdl%3D1